Our mission is to utilize basic knowledge of an enzymatic function, often in combination with structural information, to create a new activity or alter selectivity of existing enzymes. Currently, we create APOBEC variants with new catalytic properties, predestinating them to use as an alternative to bisulfite treatment in determining the pattern of DNA/RNA methylation and hydroxymethylation.
Technology
Our mission
Why APOBECs?
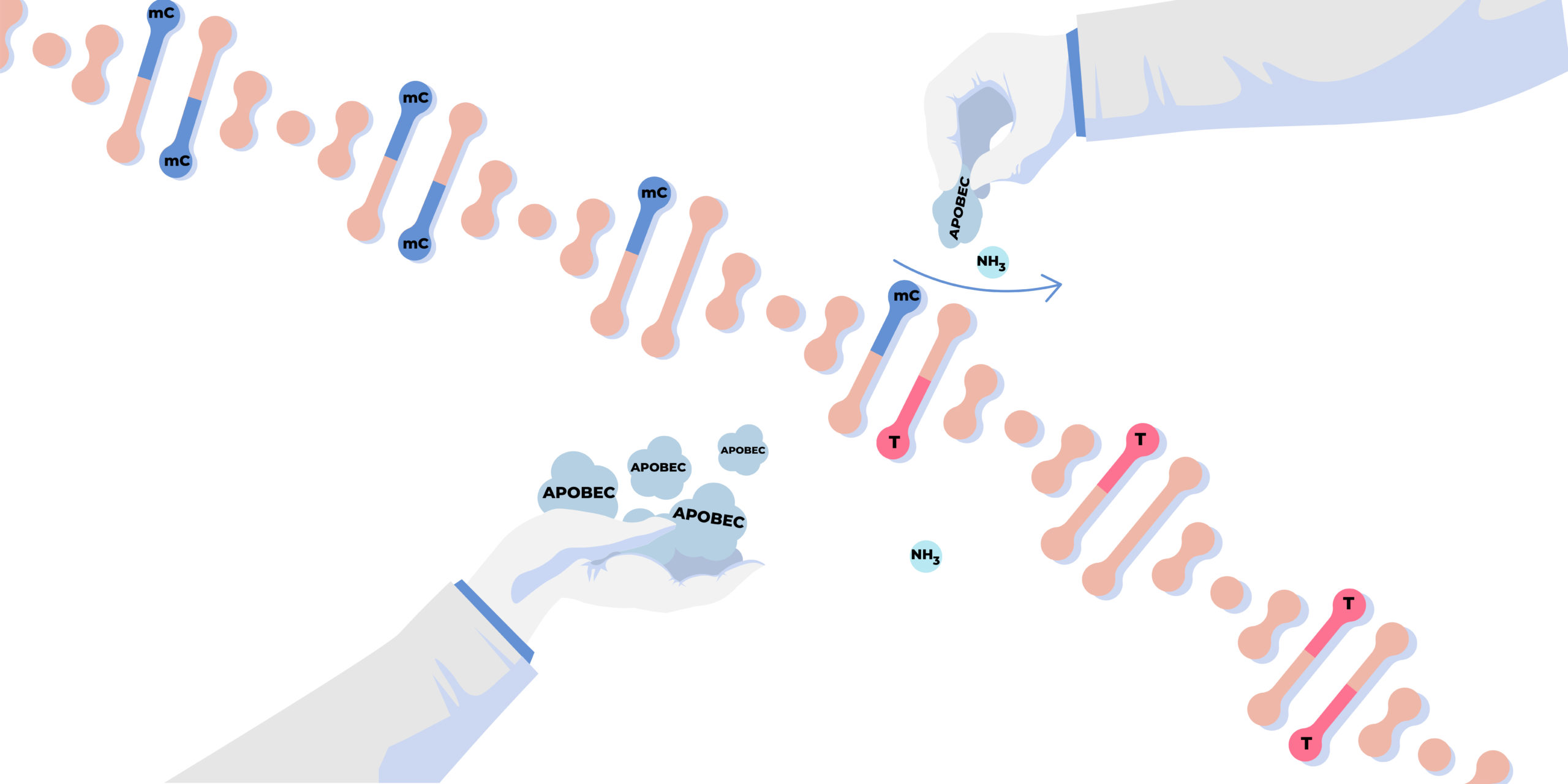
AID/APOBEC family represents a unique group of enzymes that function as DNA and RNA mutators. They can insert mutations in DNA and/or RNA as a result of their ability to deaminate cytidine to uridin. In addition to the canonical deamination of cytidine, four APOBEC members (AID, APOBEC3A, APOBEC3B, and APOBEC3H) have been demonstrated to deaminate methylated cytidines. Characteristics of the family – high similarity of structures with simultaneous specialization in recognizing different substrates – determine a high plasticity of these proteins and a wide range of possibilities for structure-guided engineering. Moreover one or a few amino acid changes are often sufficient to modulate their enzymatic activity or selectivity, while maintaining the overall protein architecture. Consequently a number of potential practical applications have been already proposed for AID/APOBECs, e.g. in base editing.
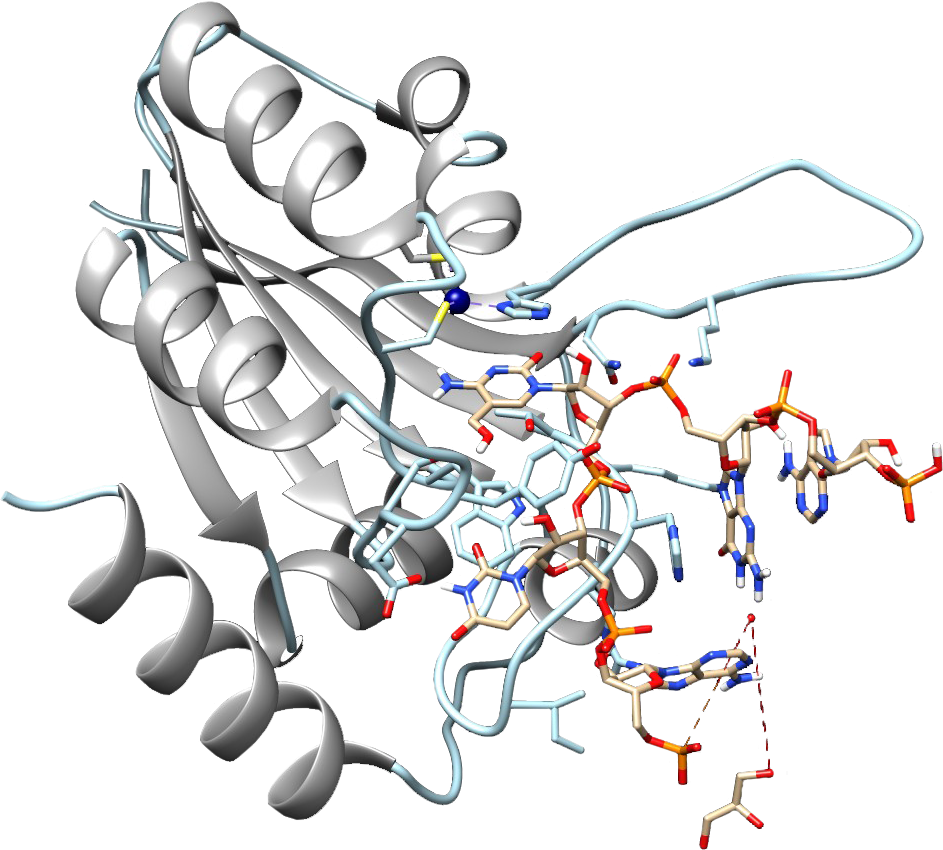
About the project
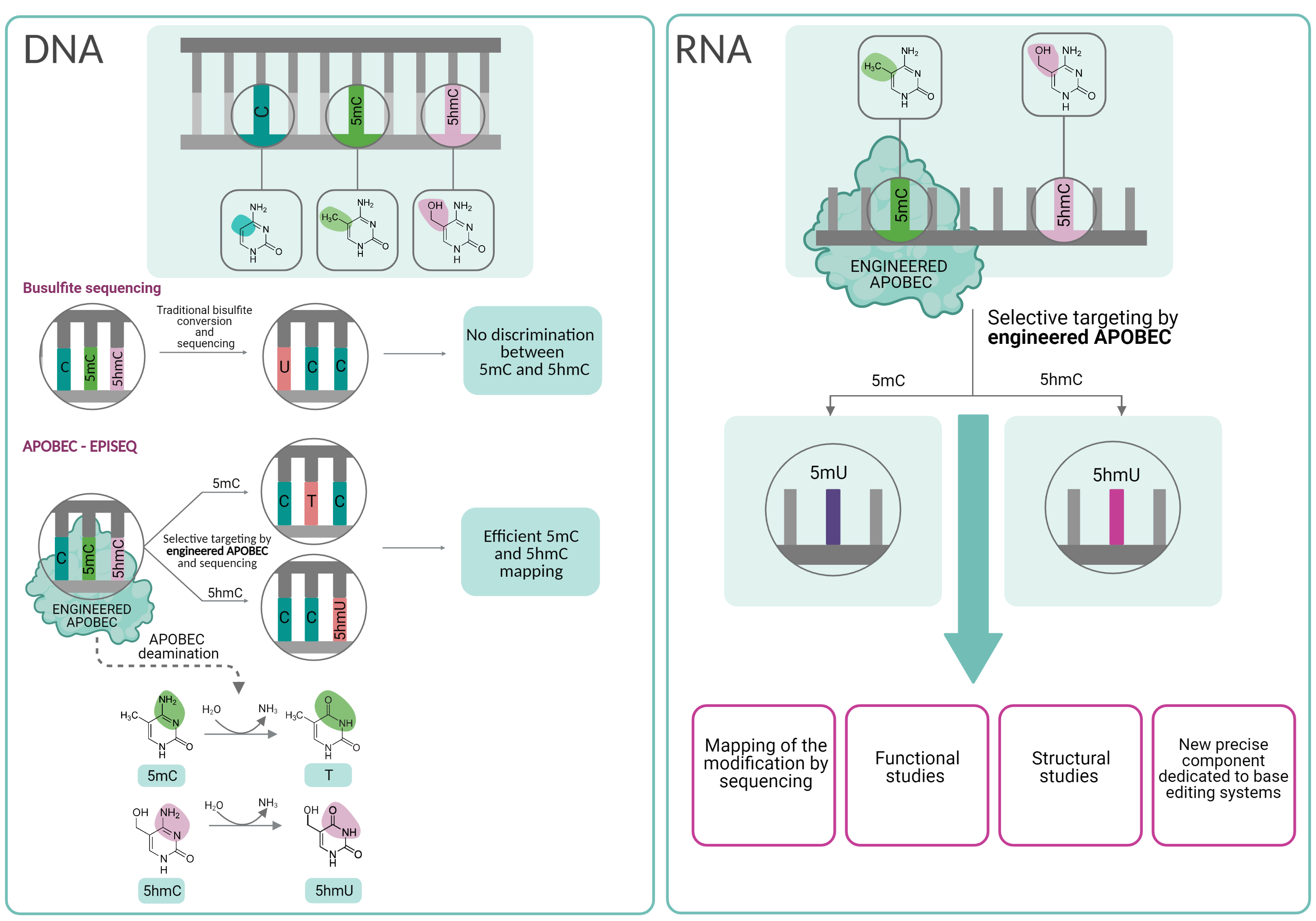
The aim of the project is to provide an innovative method and products with a global potential. The proposed products have the form of engineered APOBEC enzymes which exhibit selective deaminase activity on 5-methylcytosine (5mC) and 5-hydroxymethylcytosine (5hmC). The enzymatic properties of AID/APOBEC proteins are unique and make the enzymes suitable for the application in cytosine modifications mapping in nucleic acids. The currently used method, bisulfite sequencing, is not able to distinguish between 5mC and 5hmC. Consequently, our knowledge of the biological role of both modifications is limited. Moreover, the use of bisulfite conversion on RNA is not optimal due to the problem with the material degradation. The method proposed in the project – Apobec-EpiSeq – overcomes technological obstacles mentioned above and demonstrates the applicability of these enzymes as promising tools in molecular biology. We have already developed a variant of hAID protein which exhibits substrate selectivity for 5mC in DNA. As part of this project, we are planning to demonstrate the applicability of this enzyme and to engineer new ones that exhibit selective deaminase activity on 5mC and 5hmC in RNA.
Objectives:
- To engineer AID/APOBEC enzymes which exhibit selective deaminase activity on 5mC and 5hmC in DNA and RNA
- To prove their appllicability in mapping of cytosine modifications
- Validation and benchmarking of the proposed methods
- Adaptation of the technology for commercial applications
Our Science
We develop efficient and accurate editors to selectively alter particular modifications in DNA and RNA molecules. Check out the diagram below to learn more about the key principles of our technology.
Our research combine imagination with the rigors of applied sciences to create novel biotechnology tools. Click here to learn more about the proposed applications.
Publications ans IP rights supporting the project
Mutations in human AID differentially affect its ability to deaminate cytidine and 5-methylcytidine in ssDNA substrates in vitro
Lucyna Budzko, Paulina Jackowiak, Karol Kamel, Joanna Sarzynska, Janusz M. Bujnicki, Marek Figlerowicz; Scientific Reports 7, 2017 doi:10.1038/s41598-017-03936-x
Production of an active human AID enzyme in a bacterial system
Lucyna Budzko, Paulina Jackowiak, Marek Figlerowicz; BioTechnologia vol. 97(3), 2016
European Patent no EP2870241
Polish Patent no PL225333
United States Patent Application no US 14/384,665
International Patent Application under the PCT no PCT/PL2014/000003
Lucyna Budzko, Paulina Jackowiak, Marek Figlerowicz „Modified gene of human activation-induced cytidine deaminase (AID), a method of modification of human AID gene, a composition showing AID activity, a method of preparation of such composition in a bacterial system and use of the composition in the analyses of DNA/RNA amination/deamination and/or methylation/demethylation”

This project has received funding from the National Centre for Research and Development under the LIDERIX Programme

„Mapping of cytosine modifications in nucleic acids using AID/APOBEC enzymes”; Project no: LIDER/30/0111/L-9/17/NCBR/2018