We develop efficient and accurate editors to selectively alter particular modifications in the genome and in RNA molecules. The selective editing of modified nucleotides can find applications in epigenomics, epitranscriptomics, and CRISPR–Cas gene editing technologies.
Applications
EPIGENOMICS:
5mC and 5hmC Sequencing
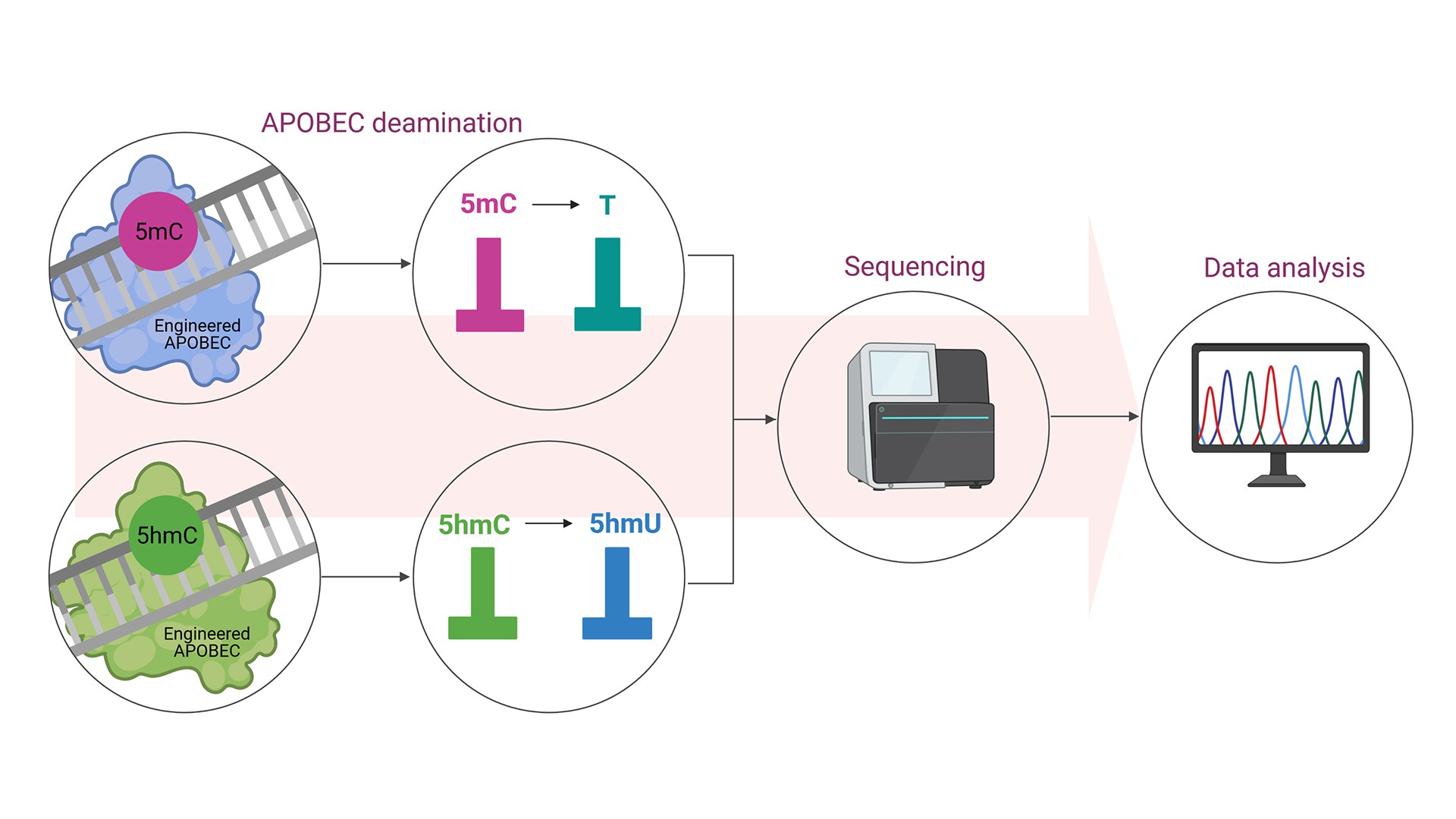
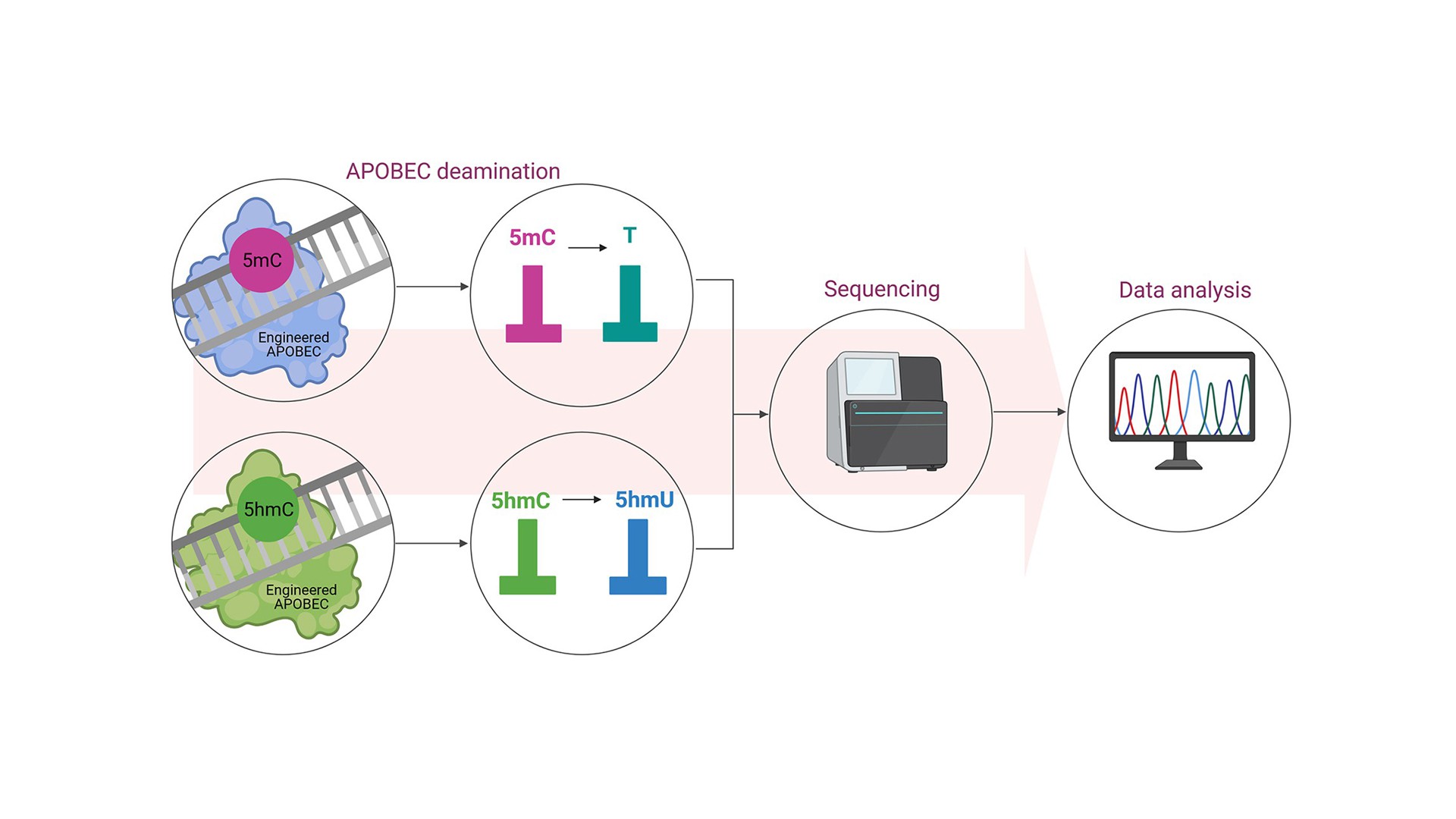
Engineered APOBECs in a combination with whole genome sequencing extend the information content of DNA sequencing by enabling two additional letters to be read: methylcytosine (5mC) and hydroxymethylcytosine (5hmC) – two the most important epigenetic marks. Importantly, this information can be captured at single base resolution.
GENE-EDITING TECHNOLOGY:
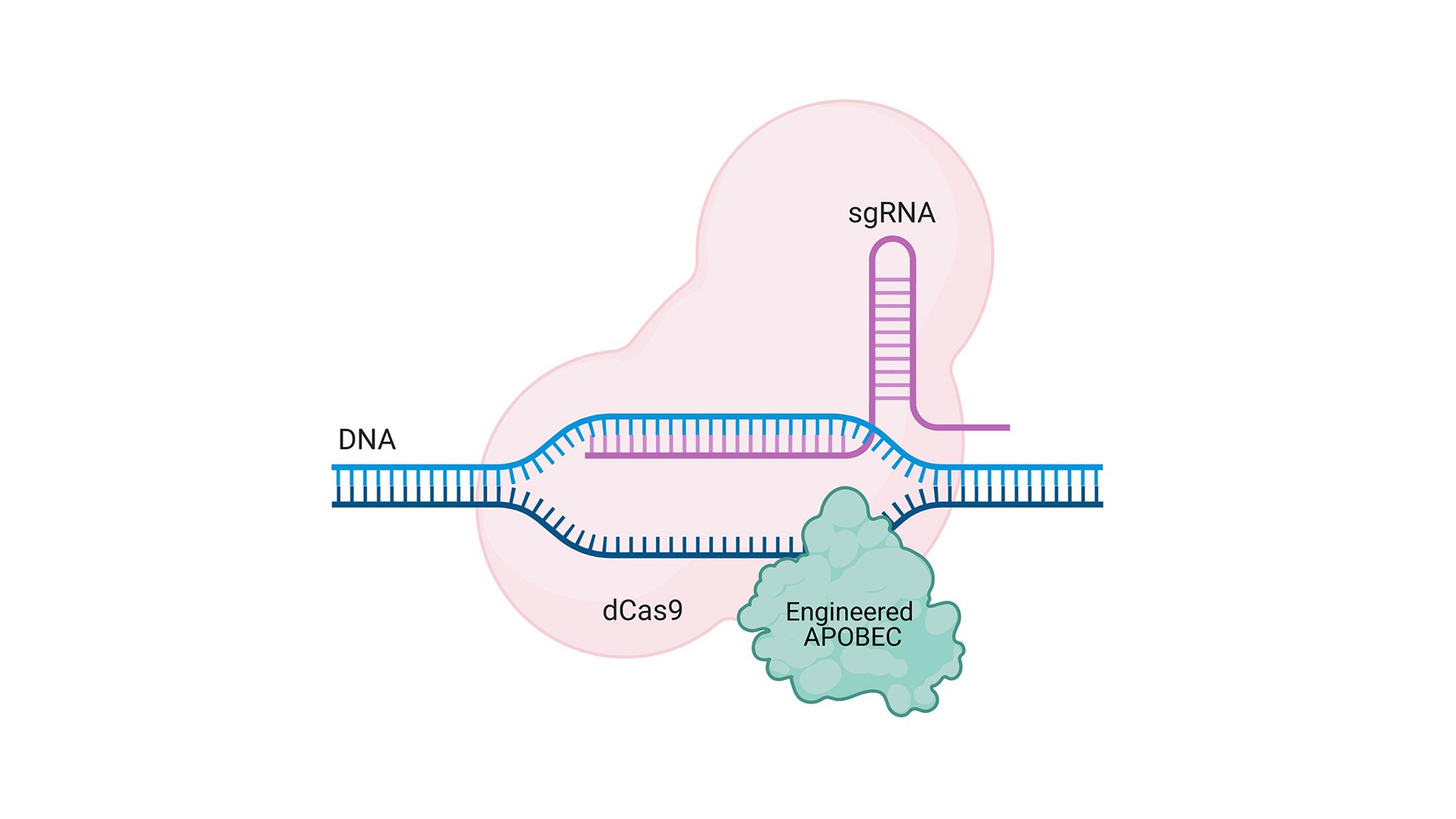
A new class of Base Editors
BASE Editing is a next-generation genome-editing tool that generates point mutations directly in cellular DNA or RNA. The ability to alter a single letter in the genetic code can correct disease-causing point mutations or create a desirable genetic variation. Currently, there are no Base Editing systems targeting modified nucleotides such as 5mC or 5hmC…
EPITRANSCRIPTOMICS:
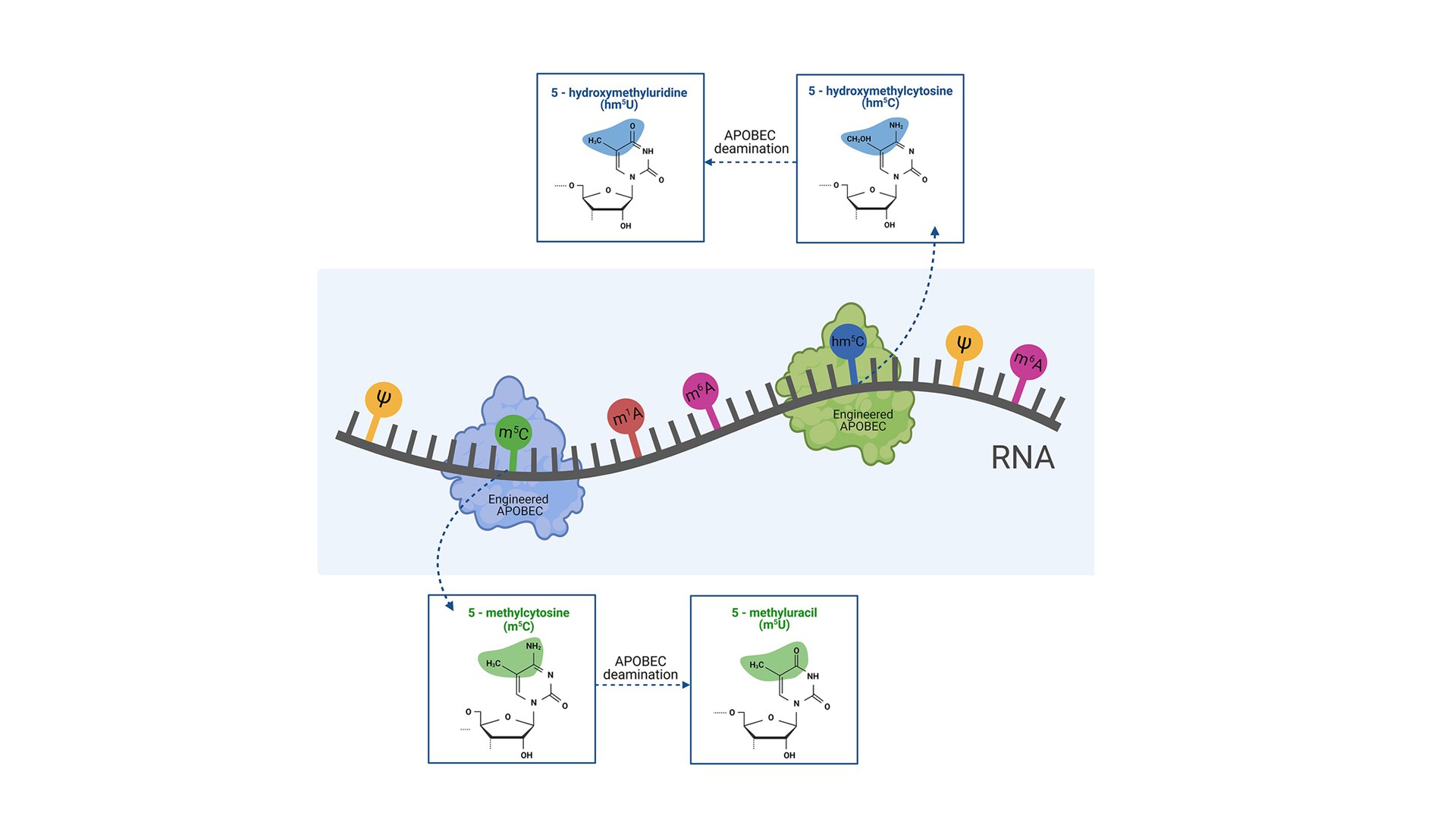
RNA modification research
Engineered APOBECs, as RNA-editing enzymes, target two common RNA modifications: methylcytosine (m5C) and hydroxymethylcytosine (hm5C), enabling their pioneering detection and expending research of their biological role. Importantly, there are no tools available on the market, to study and manipulate RNA modifications.
5mC and 5hmC Sequencing
Epigenetic changes, such as DNA methylation and hydroxymethylation, are a heritable layer of genetic information that are not attributable to changes in the primary DNA sequence. Epigenetic modifications are essential for gene regulation and expression, and thereby underlie the development, regulation, and maintenance of the normal cell. Lifestyle, nutrition, and environmental factors can all lead to epigenetic changes. Epigenetic dysregulations are associated with several diseases, including cancer. 5mC is the most common modified base in the genome, and hence it is considered as the fifth genetic code (in addition to the canonical A, G, C, T). 5hmC is abundant, enzymatically derived modification of 5mC in mammalian DNA, which levels vary significantly depending on the cell type and increase with age. The most common epigenetic analysis – bisulfite sequencing, is not able to distinguish between 5mC and 5hmC. Consequently, our knowledge of these two modifications is still limited. Engineered APOBECs overcome this technological obstacle through selective deamination of 5mC or 5hmC. Consequently these engineered editors may extend the possibilities of currently used sequencing methods.
A new class of Base Editors
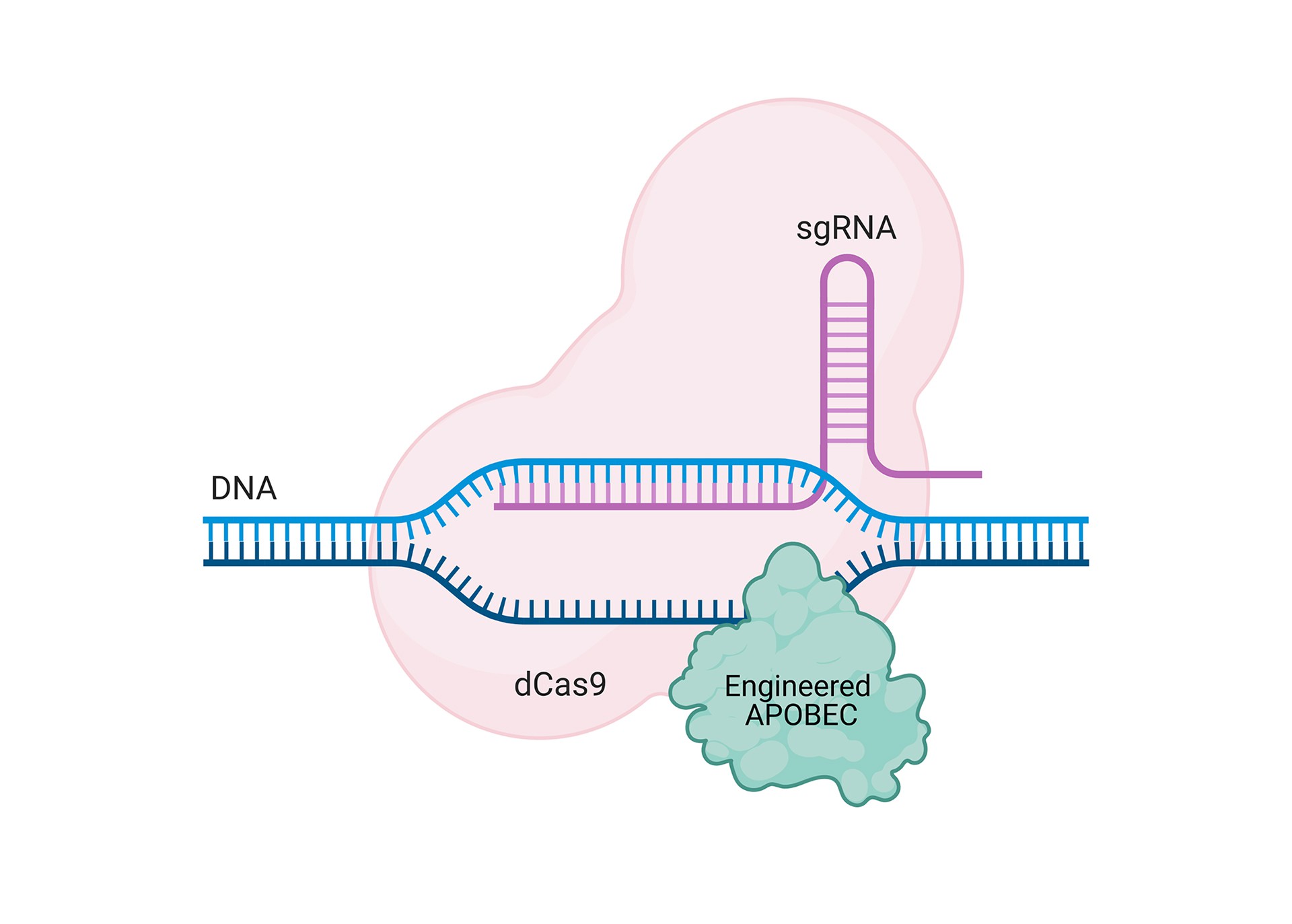
CRISPR-Cas gene-editing technology is a genetic engineering technique by which the genomes of living organisms may be modified. This method has been first published in 2012 by Jennifer Doudna and Emmanuelle Charpentier, Nobel Laureates in 2020. Since 2012 this technology highly developed, revealing an unexpected link between APOBECs and CRISPR-Cas in so called BASE Editing system. BASE Editing is a next-generation genome-editing tool that uses components of the CRISPR-Cas technology (a catalytically impaired Cas9 variant) together with APOBEC enzymes (or other deaminases) to generate point mutations directly in cellular DNA or RNA. Komor and coworkers created the first cytosine Base Editor by coupling a cytidine deaminase with the inactive dCas9 (Komor et al., 2016). This fusion convert cytosine to uracil, without cutting DNA, and uracil is then subsequently converted to thymine through DNA replication or repair. Base editors have the ability to change just a single letter in the genetic code, what can mean the difference between health and disease. Therefore, variants of this system are widely developed in scientific laboratories around the world. Importantly, there are no Base Editing system targeting modified nucleotides such as 5mC or 5hmC. We are devoting our efforts to fill this gap, and provide the new editing component to the technology.
RNA modification research
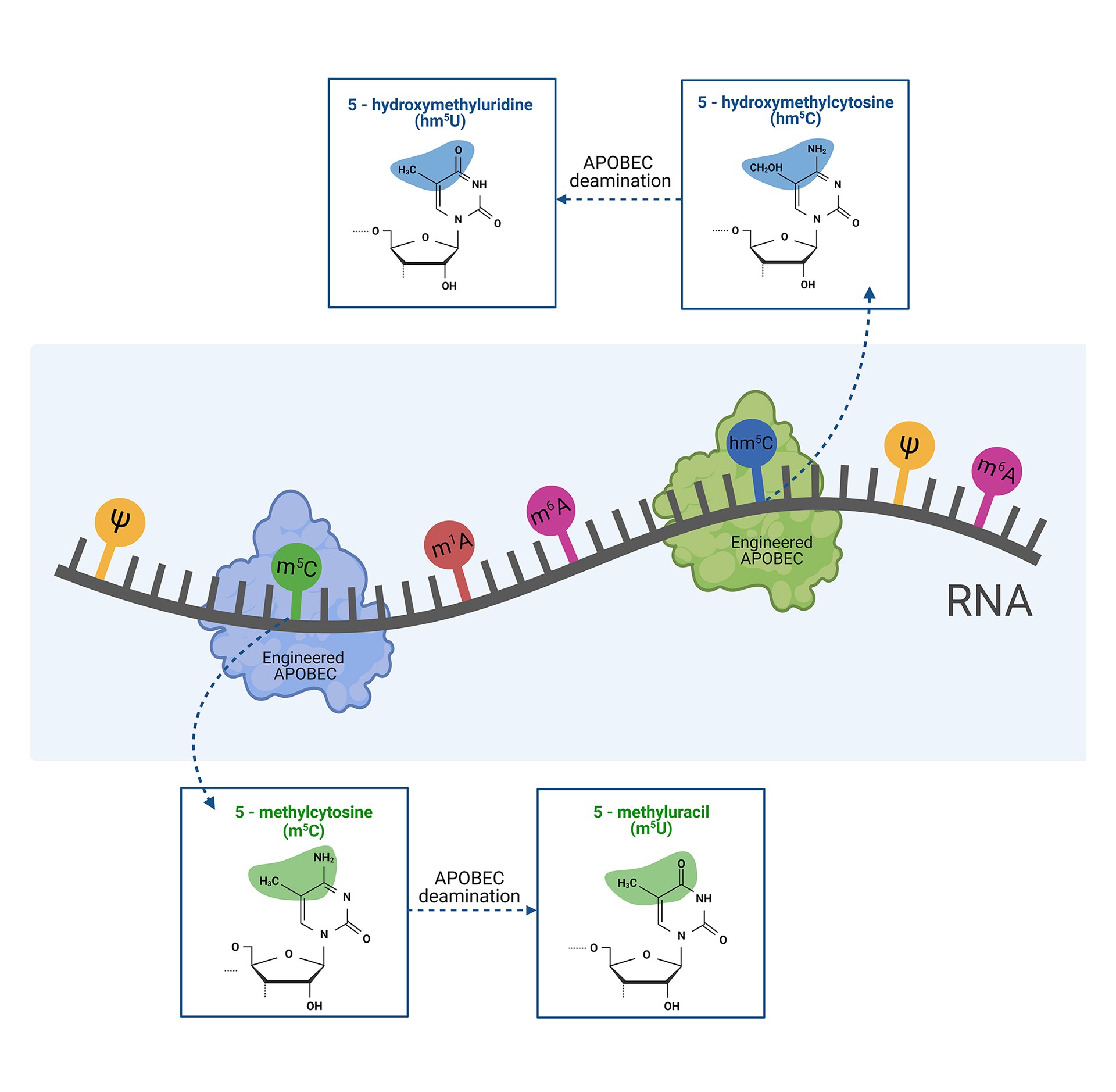
The epitranscriptome is just starting to emerge, understood as RNA nucleotide modifications that regulate and alter the activity of RNA molecules. Currently, more than 150 different RNA modifications have been identified, occurring in organisms ranging from archaea and bacteria, to eukaryotes. For most of them, we still have limited understanding of their biological role. Three members of APOBEC family (APOBEC1, APOBEC3A, APOBEC3G) have been demonstrated to deaminate cytidines in RNA. A high plasticity of the protein family and the proven activity on modified nucleotides in DNA inspired us to engineer APOBEC variants with selective deaminase activity on two common RNA modifications: methylcytosine (m5C) and hydroxymethylcytosine (hm5C). The engineered activity has two effects: (1) it erases m5C or hm5C modifications (abundant in RNA molecules), and (2) generates m5U and hm5U modifications (relatively rare in RNA), enabling functional studies of four modifications in total. We aim to provide novel molecular biology tools dedicated to a pioneering detection of these RNA modifications, studies of their biological role, and their influence on RNA structure. Targeting RNA modifications could also be a promising therapeutic strategy. By editing RNA, rather than the genome itself, effects on protein function become temporary and regulatable. RNA editing could be suitable for a therapy of non-hereditary disorders, in contrast to the genome editing, mostly suitable for rare and very specific genetic disorders.
We are open to new ideas regarding potential applications of our technology in an area of your interest.
We can offer full length, human APOBEC3A recombinant protein (both wild type and selected mutants) with guaranteed protein quantity, high purity (>95%), and confirmed deamination activity.
Please feel free to contact us if you need any further information.

This project has received funding from the National Centre for Research and Development under the LIDERIX Programme

„Mapping of cytosine modifications in nucleic acids using AID/APOBEC enzymes”; Project no: LIDER/30/0111/L-9/17/NCBR/2018